Permeability prediction loglan using Genetic Algorithms.
This logan:
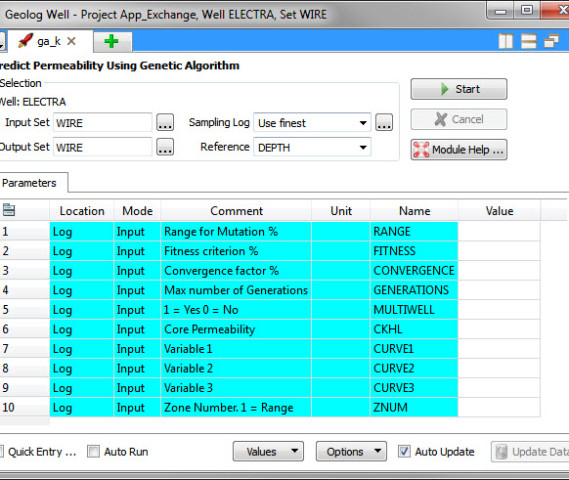
- ga_k – evolves (calibrates) an equation that relates core permeability to the logs.
Input curves: The Genetic Algorithms loglan (ga_k) requires up to three input logs. These could be DTP, DTS, RHOB, GR, RT and NPHI.
Input constants: None
Input options: RANGE, GENERATIONS and MULTIWELL The default input constants are 0.1, 20000 and 0. Change the MULTIWELL from 0 to 1 if the calibration is to be made over several wells.
Useful Notes:
- Before running plot all variables against each other on a simple Xplot. Check that there are no points off the Xplot.
- Delete all rogue points.
- Core data must be precisely depth matched before running the correlation Logan or any GAFL. It is acceptable to manually shift data at formation shoulders.
- If you use the gamma-ray it must be normalised for multiple well processing.
- When normalising the GR ensure that less than 5% of the points are truncated at 1. Otherwise you will lose GR information.
- Change the RANGE and GENERATIONS to see if you can improve your results.
- Calibration file created is called ga_k_out.dat and is located in the SPECS folder.
- For Gas use numerical zones.
- Do not use any curve that exists in a different range in the prediction wells compared to the calibration wells. SW, NPHI and DEPTH are important curves to check for this.
- If you want to test the calibration quickly don’t change the number of iterations. Select a shorter interval and set the fitness at 0.1.
- For the final pass set: Fitness Criterion = 0.0001, Initial Mutation Range = 0.2, Convergence = 0.9, Maximum Number of Iteration = 50000.
- Check that the 3rd column in the Geolog screen readout (FITNESS) is a greater than zero and decreases every 1000 generations. Check that the last column is the expected number of data levels.
- Inspect the calibration data file is being updated and the values look reasonable.